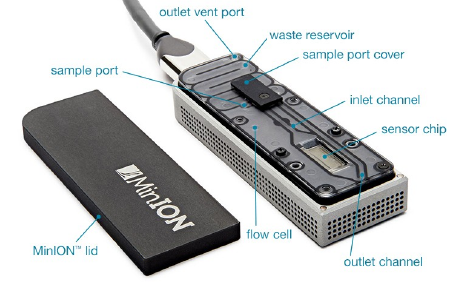
MinION (Oxford Nanopore)
Contents
Introduction
Entirely unique sequencing method, where the flowcell is inserted into a USB container, and from there, plugged into a computer.
Due to its small size in comparison with Illumina, IonTorrent and PacBio, this sequencing tool is eminently suited to field work.
Overview
Reputed advantages
- flowcell pores good for several runs, until they die out, which they may do at different times.
- Reads an be quite long ... 100kb is possible.
Shortcomings
- Computer, usually a laptop, needs to be continually connected to internet, and to be in high workload mode (no economy nor sleep mode allowed).
- accuracy at least an order of magnitude worse than Illumina (~90% vs >99%)
Software Round-up
The software required can be split into two groups of programs:
- Sequencing generation
- MinKNOW, for control of MinION device & run parameters
- Metrichor, for cloud basecalling of event data
- Chronolapse a screen image grabber for record keeping
- TeamViewer, for remote control of MinION computer
- MinoTour, live monitoring / control of run while sequencing (a collaboration with Matt Loose of Nottingham University).
- Sequence File Analysis
- Poretools, poRe Sequence extraction and data summaries (deevloped by Nick Loman and Aaron Quinlan (latter of bedtools fame)).
Links
- Brian Naugton's blog entry 11 Oct 2016 taking stock of recent advances
- Nature paper 11 Feb 2016 describing Minion use in Ebola outbreak